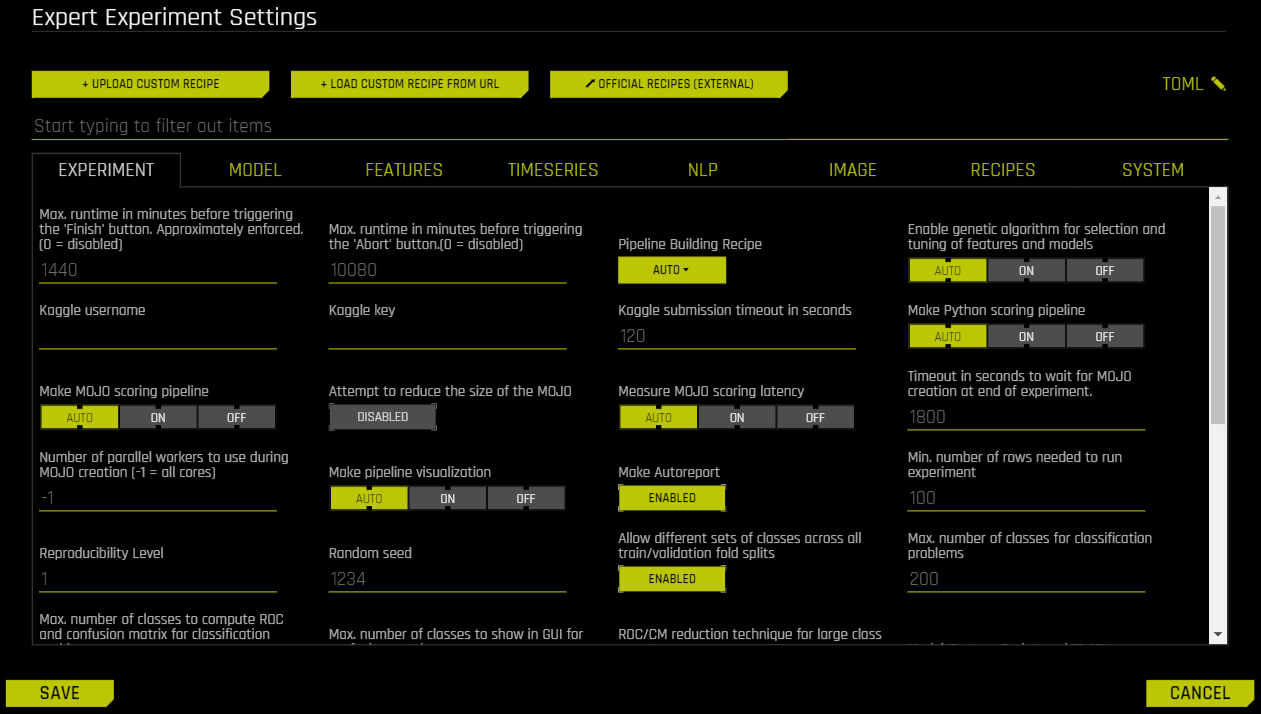
Expert Settings¶
This section describes the Expert Settings that are available when starting an experiment. Driverless AI provides a variety of options in the Expert Settings that let you customize your experiment. Use the search bar to refine the list of settings or locate a specific setting.
The default values for these options are derived from the configuration options in the config.toml file. For more information about each of these options, see the Sample config.toml File section. When a setting is changed from the default value, it is highlighted in the interface to indicate that the default value is not currently selected.
Note about Feature Brain Level: By default, the feature brain pulls in any better model regardless of the features even if the new model disabled those features. For full control over features pulled in via changes in these Expert Settings, users should set the Feature Brain Level option to 0.
- Upload Custom Recipe
- Load Custom Recipe from URL
- Official Recipes (Open Source)
- Edit the TOML Configuration
- Experiment Settings
- Max Runtime in Minutes Before Triggering the Finish Button
- Max Runtime in Minutes Before Triggering the Abort Button
- Pipeline Building Recipe
- Kaggle Username
- Kaggle Key
- Kaggle Submission Timeout in Seconds
- Make Python Scoring Pipeline
- Make MOJO Scoring Pipeline
- Attempt to Reduce the Size of the MOJO
- Measure MOJO Scoring Latency
- Timeout in Seconds to Wait for MOJO Creation at End of Experiment
- Number of Parallel Workers to Use During MOJO Creation
- Make Pipeline Visualization
- Make AutoDoc
- Min Number of Rows Needed to Run an Experiment
- Reproducibility Level
- Random Seed
- Allow Different Sets of Classes Across All Train/Validation Fold Splits
- Max Number of Classes for Classification Problems
- Max Number of Classes to Compute ROC and Confusion Matrix for Classification Problems
- Max Number of Classes to Show in GUI for Confusion Matrix
- ROC/CM Reduction Technique for Large Class Counts
- Model/Feature Brain Level
- Feature Brain Save Every Which Iteration
- Feature Brain Restart from Which Iteration
- Feature Brain Refit Uses Same Best Individual
- Feature Brain Adds Features with New Columns Even During Retraining of Final Model
- Restart-Refit Use Default Model Settings If Model Switches
- Min DAI Iterations
- Select Target Transformation of the Target for Regression Problems
- Enable Genetic Algorithm for Selection and Tuning of Features and Models
- Tournament Model for Genetic Algorithm
- Number of Cross-Validation Folds for Feature Evolution
- Number of Cross-Validation Folds for Final Model
- Force Only First Fold for Models
- Max Number of Rows Times Number of Columns for Feature Evolution Data Splits
- Max Number of Rows Times Number of Columns for Reducing Training Dataset
- Maximum Size of Validation Data Relative to Training Data
- Perform Stratified Sampling for Binary Classification If the Target Is More Imbalanced Than This
- Add to config.toml via toml String
- Model Settings
- Constant Models
- Decision Tree Models
- GLM Models
- XGBoost GBM Models
- LightGBM Models
- XGBoost Dart Models
- Enable RAPIDS-cuDF extensions to XGBoost GBM/Dart
- Enable Dask_cuDF (multi-GPU) XGBoost GBM
- Enable Dask_cuDF (multi-GPU) XGBoost Dart
- Enable Dask (multi-node) LightGBM
- Enable Dask (multi-node/multi-GPU) hyperparameter search
- Number of trials for hyperparameter optimization within XGBoost GBM/Dart
- Number of trials for rapids-cudf hyperparameter optimization within XGBoost GBM/Dart for final model only
- TensorFlow Models
- PyTorch Models
- FTRL Models
- RuleFit Models
- Zero-Inflated Models
- LightGBM Boosting Types
- LightGBM Categorical Support
- LightGBM CUDA Support
- Whether to Show Constant Models in Iteration Panel
- Parameters for TensorFlow
- Max Number of Trees/Iterations
- n_estimators List to Sample From for Model Mutations for Models That Do Not Use Early Stopping
- Minimum Learning Rate for Final Ensemble GBM Models
- Maximum Learning Rate for Final Ensemble GBM Models
- Reduction Factor for Max Number of Trees/Iterations During Feature Evolution
- Minimum Learning Rate for Feature Engineering GBM Models
- Max Learning Rate for Tree Models
- Max Number of Epochs for TensorFlow/FTRL
- Max Tree Depth
- Max max_bin for Tree Features
- Max Number of Rules for RuleFit
- Ensemble Level for Final Modeling Pipeline
- Cross-Validate Single Final Model
- Number of Models During Tuning Phase
- Sampling Method for Imbalanced Binary Classification Problems
- Threshold for Minimum Number of Rows in Original Training Data to Allow Imbalanced Sampling
- Ratio of Majority to Minority Class for Imbalanced Binary Classification to Trigger Special Sampling Techniques (if Enabled)
- Ratio of Majority to Minority Class for Heavily Imbalanced Binary Classification to Only Enable Special Sampling Techniques (if Enabled)
- Number of Bags for Sampling Methods for Imbalanced Binary Classification (if Enabled)
- Hard Limit on Number of Bags for Sampling Methods for Imbalanced Binary Classification
- Hard Limit on Number of Bags for Sampling Methods for Imbalanced Binary Classification During Feature Evolution Phase
- Max Size of Data Sampled During Imbalanced Sampling
- Target Fraction of Minority Class After Applying Under/Over-Sampling Techniques
- Max Number of Automatic FTRL Interactions Terms for 2nd, 3rd, 4th order interactions terms (Each)
- Whether to Enable Bootstrap Sampling for Validation and Test Scores
- For Classification Problems with This Many Classes, Default to TensorFlow
- Compute Prediction Intervals
- Confidence Level for Prediction Intervals
- Features Settings
- Feature Engineering Effort
- Data Distribution Shift Detection
- Data Distribution Shift Detection Drop of Features
- Max Allowed Feature Shift (AUC) Before Dropping Feature
- Data Leakage Detection
- Data Leakage Detection Dropping AUC/R2 Threshold
- Max Rows Times Columns for Leakage
- Report Permutation Importance on Original Features
- Maximum Number of Rows to Perform Permutation-Based Feature Selection
- Max Number of Original Features Used
- Max Number of Original Non-Numeric Features
- Max Number of Original Features Used for FS Individual
- Number of Original Numeric Features to Trigger Feature Selection Model Type
- Number of Original Non-Numeric Features to Trigger Feature Selection Model Type
- Max Allowed Fraction of Uniques for Integer and Categorical Columns
- Allow Treating Numerical as Categorical
- Max Number of Unique Values for Int/Float to be Categoricals
- Max Number of Engineered Features
- Max Number of Genes
- Limit Features by Interpretability
- Threshold for Interpretability Above Which to Enable Automatic Monotonicity Constraints for Tree Models
- Correlation Beyond Which to Trigger Monotonicity Constraints (if enabled)
- Control amount of logging when calculating automatic monotonicity constraints (if enabled)
- Whether to drop features that have no monotonicity constraint applied (e.g., due to low correlation with target)
- Manual Override for Monotonicity Constraints
- Max Feature Interaction Depth
- Fixed Feature Interaction Depth
- Enable Target Encoding
- Enable Outer CV for Target Encoding
- Enable Lexicographical Label Encoding
- Enable Isolation Forest Anomaly Score Encoding
- Enable One HotEncoding
- Number of Estimators for Isolation Forest Encoding
- Drop Constant Columns
- Drop ID Columns
- Don’t Drop Any Columns
- Features to Drop
- Features to Group By
- Sample from Features to Group By
- Aggregation Functions (Non-Time-Series) for Group By Operations
- Number of Folds to Obtain Aggregation When Grouping
- Type of Mutation Strategy
- Enable Detailed Scored Features Info
- Enable Detailed Logs for Timing and Types of Features Produced
- Compute Correlation Matrix
- Required GINI Relative Improvement for Interactions
- Number of Transformed Interactions to Make
- Whether to enable RAPIDS cuML GPU transformers (no mojo)
- Time Series Settings
- Time-Series Lag-Based Recipe
- Larger Validation Splits for Lag-Based Recipe
- Fixed-size train timespan across splits
- Custom Validation Splits for Time-Series Experiments
- Timeout in Seconds for Time-Series Properties Detection in UI
- Generate Holiday Features
- List of Countries for Which to Look up Holiday Calendar and to Generate Is-Holiday Features For
- Time-Series Lags Override
- Lags override for features that are not known ahead of time
- Lags override for features that are known ahead of time
- Smallest Considered Lag Size
- Enable Feature Engineering from Time Column
- Allow Integer Time Column as Numeric Feature
- Allowed Date and Date-Time Transformations
- Auto filtering of date and date-time transformations
- Consider Time Groups Columns as Standalone Features
- Which TGC Feature Types to Consider as Standalone Features
- Enable Time Unaware Transformers
- Always Group by All Time Groups Columns for Creating Lag Features
- Generate Time-Series Holdout Predictions
- Number of Time-Based Splits for Internal Model Validation
- Maximum Overlap Between Two Time-Based Splits
- Maximum Number of Splits Used for Creating Final Time-Series Model’s Holdout Predictions
- Whether to Speed up Calculation of Time-Series Holdout Predictions
- Whether to Speed up Calculation of Shapley Values for Time-Series Holdout Predictions
- Generate Shapley Values for Time-Series Holdout Predictions at the Time of Experiment
- Lower Limit on Interpretability Setting for Time-Series Experiments (Implicitly Enforced)
- Dropout Mode for Lag Features
- Probability to Create Non-Target Lag Features
- Method to Create Rolling Test Set Predictions
- Fast TTA for GA Validation
- Probability for New Time-Series Transformers to Use Default Lags
- Probability of Exploring Interaction-Based Lag Transformers
- Probability of Exploring Aggregation-Based Lag Transformers
- Time Series Centering or Detrending Transformation
- Custom Bounds for SEIRD Epidemic Model Parameters
- Which SEIRD Model Component the Target Column Corresponds To
- Time Series Lag-Based Target Transformation
- Lag Size Used for Time Series Target Transformation
- NLP Settings
- Max TensorFlow Epochs for NLP
- Accuracy Above Enable TensorFlow NLP by Default for All Models
- Enable Word-Based CNN TensorFlow Models for NLP
- Enable Word-Based BiGRU TensorFlow Models for NLP
- Enable Character-Based CNN TensorFlow Models for NLP
- Enable PyTorch Models for NLP (Experimental)
- Select Which Pretrained PyTorch NLP Models to Use
- Number of Epochs for Fine-Tuning of PyTorch NLP Models
- Batch Size for PyTorch NLP Models
- Maximum Sequence Length for PyTorch NLP Models
- Path to Pretrained PyTorch NLP Models
- Path to Pretrained Embeddings for TensorFlow NLP Models
- Allow Training of Unfrozen Pretrained Embeddings
- Fraction of Text Columns Out of All Features to be Considered a Text-Dominanted Problem
- Fraction of Text per All Transformers to Trigger That Text Dominated
- Threshold for String Columns to be Treated as Text
- Max Size of the Vocabulary for Text Transformers
- Image Settings
- Enable Image Transformer for Processing of Image Data
- Supported ImageNet Pretrained Architectures for Image Transformer
- Dimensionality of Feature Space Created by Image Transformer
- Enable Fine-Tuning of the Pretrained Models Used for the Image Transformer
- Number of Epochs for Fine-Tuning Used for the Image Transformer
- List of Augmentations for Fine-Tuning Used for the Image Transformer
- Batch Size for the Image Transformer
- Image Download Timeout in Seconds
- Maximum Allowed Fraction of Missing Values for Image Column
- Minimum Fraction of Images That Need to Be of Valid Types for Image Column to Be Used
- Enable GPU(s) for Faster Transformations With the Image Transformer
- Recipes Settings
- Include Specific Transformers
- Include Specific Preprocessing Transformers
- Number of Pipeline Layers
- Include Specific Models
- Include Specific Scorers
- Scorer to Optimize Threshold to Be Used in Other Confusion-Matrix Based Scorers (For Binary Classification)
- Include Specific Data Recipes During Experiment
- Probability to Add Transformers
- Probability to Add Best Shared Transformers
- Probability to Prune Transformers
- Probability to Mutate Model Parameters
- Probability to Prune Weak Features
- Timeout in Minutes for Testing Acceptance of Each Recipe
- Whether to Skip Failures of Transformers
- Whether to Skip Failures of Models
- Level to Log for Skipped Failures
- System Settings
- Exclusive level of access to node resources
- Number of Cores to Use
- Maximum Number of Cores to Use for Model Fit
- If full dask cluster is enabled, use full cluster
- Maximum Number of Cores to Use for Model Predict
- Maximum Number of Cores to Use for Model Transform and Predict When Doing MLI, AutoDoc
- Tuning Workers per Batch for CPU
- Number of Workers for CPU Training
- #GPUs/Experiment
- Num Cores/GPU
- #GPUs/Model
- Num. of GPUs for Isolated Prediction/Transform
- Max Number of Threads to Use for datatable and OpenBLAS for Munging and Model Training
- Max Number of Threads to Use for datatable Read and Write of Files
- Max Number of Threads to Use for datatable Stats and OpenBLAS
- GPU Starting ID
- Enable Detailed Traces
- Enable Debug Log Level
- Enable Logging of System Information for Each Experiment
- AutoDoc Settings
- Make AutoDoc
- AutoDoc Name
- AutoDoc Template Location
- AutoDoc File Output Type
- AutoDoc SubTemplate Type
- Confusion Matrix Max Number of Classes
- Number of Top Features to Document
- Minimum Relative Feature Importance Threshold
- Permutation Feature Importance
- Number of Permutations for Feature Importance
- Feature Importance Scorer
- PDP Max Number of Rows
- PDP Max Runtime in Seconds
- PDP Out of Range
- ICE Number of Rows
- Population Stability Index
- Population Stability Index Number of Quantiles
- Prediction Statistics
- Prediction Statistics Number of Quantiles
- Response Rates Plot
- Response Rates Plot Number of Quantiles
- Show GINI Plot
- Enable Shapley Values
- Number of Features in Data Summary Table
- List All Config Settings
- Keras Model Architecture Summary Line Length
- NLP/Image Transformer Architecture Max Lines
- Appendix NLP/Image Transformer Architecture
- Full GLM Coefficients Table in the Appendix
- GLM Coefficient Tables Number of Models
- GLM Coefficient Tables Number of Folds Per Model
- GLM Coefficient Tables Number of Coefficients
- GLM Coefficient Tables Number of Classes
- Number of Histograms to Show